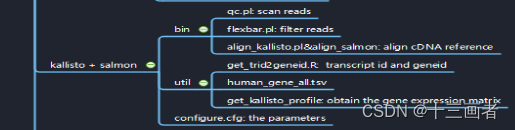
RNA-seq pipeline through kallisto or salmon
kallisto 和salmon相比含有hisat2和STAR等软的RNA-seq流程而言,速度更快,这是因为该软件基于转录组序列reference(也即是cDNA序列)并且基于k mer比对原理。通常如果想研究RNA-seq过程基因发生了何种变化且不需知道新转录本,可以直接使用kallisto或salmon获取转录本的丰度信息。
install software
conda install -c bioconda kallisto salmon -y
过滤原始 fastq files
任何RNA-seq流程都需要对原始fastq reads文件进行质控,质控包括去除接头、barcode等额外引入的序列,当然也包括舍弃低质量的reads。这类软件有,cutadapt、flexbar和trimmomatic等。
download reference
转录本序列可以从多个网站下载,这里我们下载ENSEMBL数据库的小鼠cDNA数据。
wget ftp://ftp.ensembl.org/pub/release-100/fasta/mus_musculus/cdna/Mus_musculus.GRCm38.cdna.all.fa
build index
kallisto index -i kallisto_index/Mus_musculus.GRCm38 Mus_musculus.GRCm38.cdna.all.fa
salmon index -i salmon_index/Mus_musculus.GRCm38 -t Mus_musculus.GRCm38.cdna.all.fa -p 20
生成批量运行的脚本
准备文件:samples.path.tsv(原始数据文件)
| SampleID | LaneID | Path |
|---|---|---|
| HF_NC01 | HF_NC01_RNA_b2.r2 | /01.rename/HF_NC01_RNA_b2.r2.fq.gz |
| HF_NC01 | HF_NC01_RNA_b2.r1 | /01.rename/HF_NC01_RNA_b2.r1.fq.gz |
| HF_NC02 | HF_NC02_RNA_b2.r1 | /01.rename/HF_NC02_RNA_b2.r1.fq.gz |
| HF_NC02 | HF_NC02_RNA_b2.r2 | /01.rename/HF_NC02_RNA_b2.r2.fq.gz |
| HF_NC03 | HF_NC03_RNA_novo.r2 | /01.rename/HF_NC03_RNA_novo.r2.fq.gz |
| HF_NC03 | HF_NC03_RNA_novo.r1 | /01.rename/HF_NC03_RNA_novo.r1.fq.gz |
过滤文件目录:02.trim (高质量的reads文件)
撰写脚本quant_feature.py:生成批量运行shell文件
# key command
kallisto quant -i kallisto_index/Mus_musculus.GRCm38 -g gtf_file -o kallisto_quant/HF07 -b 30 -t 10 02.trim/HF07_1.fastq.gz 02.trim/HF07_2.fastq.gzsalmon quant -i salmon_index/Mus_musculus.GRCm38/ -l IU -p 10 -1 02.trim/HF07_1.fastq.gz -2 02.trim/HF07_2.fastq.gz -o salmon_quant/HF07 --numBootstraps 30
#!/usr/bin/pythonimport os
import re
import sys
import argparse as ap
from collections import defaultdictcurrent_abosulte_path = os.getcwd()def parse_arguments(args):parser = ap.ArgumentParser(description= "the initial script of pipeline")parser.add_argument('-t','--trim',metavar='<string>', type=str, help='trimmed data dir')parser.add_argument('-s','--sample',metavar='<string>', type=str, help='SampleID|LaneID|Path')parser.add_argument('-o','--output',metavar='<string>', type=str, help='the output directory')parser.add_argument('-g','--gtf',metavar='<string>', type=str, help='the genome annotation file')parser.add_argument('-i','--index',metavar='<string>', type=str, help='the index of genome')parser.add_argument('-p','--prefix',metavar='<string>', type=str, help='the prefix of output file')return parser.parse_args()def create_folder(folder):'''create folder for the result'''folder_path = os.path.join(current_abosulte_path, folder)if not os.path.exists(folder_path):os.makedirs(folder_path)return folder_pathdef get_path_file(sample):'''the connections among sampleid, laneid and path'''dict_lane = defaultdict(list)with open(sample, 'r') as f:for line in f:line = line.strip()if not line.startswith("SampleID"):group = re.split(r'\s+', line)dict_lane[group[0]].append(group[1])return dict_lanedef generate_shell(trim_folder, sample_file, outdir, gtf_file, genome_index, file_name):trim_file_folder = os.path.join(current_abosulte_path, trim_folder)output_path = create_folder(outdir)output_name = os.path.join(current_abosulte_path, file_name)sample_name = get_path_file(sample_file)output_f = open(output_name, 'w')for key in sample_name:out_sample_prefix = os.path.join(output_path, key)#target1 = [x for x in sample_name[key] if re.findall(r'r1', x)]#fq1 = os.path.join(trim_file_folder, ("_".join([target1[0], "val_1.fq.gz"])))#target2 = [x for x in sample_name[key] if re.findall(r'r2', x)]#fq2 = os.path.join(trim_file_folder, ("_".join([target2[0], "val_2.fq.gz"])))fq1 = os.path.join(trim_file_folder, ("_".join([key, "1.fastq.gz"])))fq2 = os.path.join(trim_file_folder, ("_".join([key, "2.fastq.gz"])))if os.path.isfile(fq1) and os.path.isfile(fq2):fq = " ".join([fq1, fq2])if re.findall(r'kallisto', genome_index):shell = " ".join(["kallisto quant -i", genome_index, "-g", \"gtf_file", "-o", out_sample_prefix, "-b 30 -t 10", fq])elif re.findall(r'salmon', genome_index):shell = " ".join(["salmon quant -i", genome_index, \"-l IU -p 10", \"-1", fq1, "-2", fq2, \"-o", out_sample_prefix,\"--numBootstraps 30"])output_f.write(shell + "\n")output_f.close()def main():args = parse_arguments(sys.argv)generate_shell(args.trim, args.sample, args.output, args.gtf, args.index, args.prefix)if __name__ == '__main__':main()
运行脚本: 生成RUN文件
# kallisto
python quant_feature.py -t 02.trim/ -s 46RNA.samples.path.tsv -o kallisto_quant -g /disk/share/database/Mus_musculus.GRCm38_release100/Mus_musculus.GRCm38.cdna.all.fa -i /disk/share/database/Mus_musculus.GRCm38_release100/kallisto_index/Mus_musculus.GRCm38 -p RUN.kallisto_quant.sh
# salmon
python quant_feature.py -t 02.trim/ -s 46RNA.samples.path.tsv -o salmon_quant -g /disk/share/database/Mus_musculus.GRCm38_release100/Mus_musculus.GRCm38.cdna.all.fa -i /disk/share/database/Mus_musculus.GRCm38_release100/salmon_index/Mus_musculus.GRCm38/ -p RUN.salmon_quant.sh
获取expression matrix:
撰写get_merged_table.R,从quant目录获取整合所有样本表达值的matrix文件
#!/bin/usr/R
library(dplyr)
library(argparser)
library(tximport)# parameter input
parser <- arg_parser("merge transcript abundance files") %>%add_argument("-s", "--sample",help = "the sampleID files") %>%add_argument("-d", "--dir",help = "the dir of output") %>%add_argument("-g", "--gene",help = "transcriptID into geneID") %>%add_argument("-t", "--type",help = "the type of software") %>%add_argument("-c", "--count",help = "the method of scale for featurecounts") %>%add_argument("-n", "--name",help = "the name of transcript file") %>%add_argument("-o", "--out",help = "the merged files's dir and name")args <- parse_args(parser)# prepare for function
sample <- args$s
dir <- args$d
tx2gen <- args$g
type <- args$t
scount <- args$c
name <- args$n
prefix <- args$otx2gene_table <- read.csv(tx2gen, header=T)
tx2gene <- tx2gene_table %>%dplyr::select(tx_id, gene_id, gene_name)
phen <- read.table(sample, header=T, sep="\t")
sample_name <- unique(as.character(phen$SampleID))
files <- file.path(dir, sample_name, name)
names(files) <- sample_name
result <- tximport(files,type = type,countsFromAbundance = scount,tx2gene = tx2gene,ignoreTxVersion = T)
save(result, file = prefix)
Rscript get_merged_table.R \-s samples.path.tsv \-d kallisto_quant/ \-g EnsDb.Mmusculus.v79_transcriptID2geneID.csv \-t kallisto \-c no \-n abundance.tsv \-o kallisto_quant.RDataRscript get_merged_table.R \-s samples.path.tsv \-d salmon_quant/ \-g EnsDb.Mmusculus.v79_transcriptID2geneID.csv \-t salmon \-c no \-n quant.sf \-o salmon_quant.RData
获取TPM和FPKM表达矩阵:可在本地电脑运行
library(dplyr)
library(tibble)
load("kallisto_quant.RData")
phen <- read.csv("phenotype.csv")
kallisto_count <- round(result$counts) %>% data.frame()
kallisto_length <- apply(result$length, 1, mean) %>% data.frame() %>%setNames("Length")get_profile <- function(matrix, exon_length,y=phen,occurrence=0.2, ncount=10){# matrix=count_format# exon_length=count_length # occurrence=0.2# ncount=10# filter with occurrence and ncountprf <- matrix %>% rownames_to_column("Type") %>% filter(apply(dplyr::select(., -one_of("Type")), 1, function(x){sum(x > 0)/length(x)}) > occurrence) %>%data.frame(.) %>% column_to_rownames("Type")prf <- prf[rowSums(prf) > ncount, ]# change sampleID sid <- intersect(colnames(prf), y$SampleID)phen.cln <- y %>% filter(SampleID%in%sid) %>%arrange(SampleID)prf.cln <- prf %>% dplyr::select(as.character(phen.cln$SampleID))# determine the right order between profile and phenotype for(i in 1:ncol(prf.cln)){ if (!(colnames(prf.cln)[i] == phen.cln$SampleID[i])) {stop(paste0(i, " Wrong"))}} colnames(prf.cln) <- phen.cln$SampleID_v2 # normalizate the profile using FPKM and TPMexon_length.cln <- exon_length[as.character(rownames(prf.cln)), , F]for(i in 1:nrow(exon_length.cln)){ if (!(rownames(exon_length.cln)[i] == rownames(prf.cln)[i])) {stop(paste0(i, " Wrong"))}} dat <- prf.cln/exon_length.cln$Lengthdat_FPKM <- t(t(dat)/colSums(prf.cln)) * 10^9dat_TPM <- t(t(dat)/colSums(dat)) * 10^6dat_count <- prf.cln[order(rownames(prf.cln)), ]dat_fpkm <- dat_FPKM[order(rownames(dat_FPKM)), ]dat_tpm <- dat_TPM[order(rownames(dat_TPM)), ]res <- list(count=dat_count, fpkm=dat_fpkm, tpm=dat_tpm)return(res)
}
prf_out <- function(result, type="STAR"){dir <- "../../Result/Profile/"if(!dir.exists(dir)){dir.create(dir)}count_name <- paste0(dir, type, "_filtered_counts.tsv")FPKM_name <- paste0(dir, type, "_filtered_FPKM.tsv")TPM_name <- paste0(dir, type, "_filtered_TPM.tsv") write.table(x = result$count, file = count_name, sep = '\t', quote = F,col.names = NA)write.table(x = result$fpkm, file = FPKM_name, sep = '\t', quote = F,col.names = NA)write.table(x = result$tpm, file = TPM_name, sep = '\t', quote = F,col.names = NA)
}kallisto_prf <- get_profile(kallisto_count, kallisto_length)
# output
prf_out(kallisto_prf, type = "kallisto")
Reference
-
kallisto manual
-
salmon github